Segway 3.0.5.dev20+gfbc360a Overview
Installation
With the Conda environment manager and the additional Bioconda channel, Segway can be installed with the command:
conda install segway
Alternatively without Bioconda, the following prerequisites must be installed for Segway:
You need Python 2.7, or Python 3.6 or later versions.
You need Graphical Models Toolkit (GMTK), which you can get at <https://github.com/melodi-lab/gmtk/releases>.
You need the HDF5 serial library and tools. The following packages are necessary for the OS you are running:
Ubuntu/Debian:
sudo apt-get install libhdf5-serial-dev hdf5-tools
CentOS/RHEL/Fedora:
sudo yum -y install hdf5 hdf5-devel
OpenSUSE:
sudo zypper in hdf5 hdf5-devel libhdf5
Afterwards Segway can be installed automatically with the command pip install
segway.
Note
Segway may not install with older versions of pip (< 6.0) due to some of its dependencies requiring the newer version. To upgrade your pip version run pip install pip –upgrade.
Standalone configuration
Segway can be run without any cluster system. This will automatically be
used when Segway fails to access any cluster system. You can force it by
setting the SEGWAY_CLUSTER environment variable to local. For example,
if you are using bash as your shell, you can run:
SEGWAY_CLUSTER=local segway
By default, Segway will use up to 32 concurrent processes when running in
standalone mode. To change this, set the SEGWAY_NUM_LOCAL_JOBS environment
variable to the appropriate number.
Cluster configuration
If you want to use Segway with your cluster, you will need the
drmaa>=0.4a3 Python package.
You need either Sun Grid Engine (SGE; now called Oracle Grid Engine), Platform Load Sharing Facility (LSF) and FedStage DRMAA for LSF, Slurm workload manager, or Torque/PBS/PBS Pro (experimental).
If FedStage DRMAA for LSF is installed, Segway should be ready to go on LSF out of the box.
If you are using the Slurm workload manager with versions past 18, it is recommended you install a DRMAA driver based on an updated fork since the original implementation is no longer updated or maintained. We currently test our Slurm support using https://github.com/natefoo/slurm-drmaa.
If you are using SGE, someone with cluster manager privileges on your
cluster must have Segway installed within their PYTHONPATH or
systemwide and then run python -m segway.cluster.sge_setup. This
sets up a consumable mem_requested attribute for every host on your
cluster for more efficient memory use.
The workflow
Segway accomplishes four major tasks from a single command-line. It–
generates an unsupervised segmentation model and initial parameters appropriate for this data;
trains parameters of the model starting with the initial parameters; and
identifies or annotates segments in this data with the model.
calculates posterior probability for each possible segment label at each position.
Note
The verbs “identify” and “annotate” are synonyms when using Segway. They both describe the same task and may be used interchangably.
Technical description
More specifically, Segway performs the following steps:
Acquires data in
genomedataformatGenerates an appropriate model for unsupervised segmentation (
segway.str,segway.inc) for use by GMTKGenerates appropriate initial parameters (
input.masterorinput.*.master) for use by GMTKWrites the data in a format usable by GMTK
Call GMTK to perform expectation maximization (EM) training, resulting in a parameter file (
params.params)Call GMTK to perform Viterbi decoding of the observations using the generated model and discovered parameters
Convert the GMTK Viterbi results into BED format (
segway.bed.gz) for use in a genome browser, or by Segtools <http://pmgenomics.ca/hoffmanlab/proj/segtools/>, or other toolsCall GMTK to perform posterior decoding of the observations using the generated model and discovered parameters
Convert the GMTK posterior results into bedGraph format (
posterior.seg*.bedGraph.gz) for use in a genome browser or other toolsUse a distributed computing system to parallelize all of the GMTK tasks listed above, and track and predict their resource consumption to maximize efficiency
Generate reports on the established likelihood at each round of training (
likelihood.*.tab)
The identify and posterior tasks can run simultaneously, as they depend only on the results of train, and not each other.
Data selection
Segway accepts data only in the Genomedata format. The Genomedata package includes utilities to convert from BED, wiggle, and bedGraph formats. By default, Segway uses all the continuous data tracks in a Genomedata archive. Multiple Genomedata archives can be specified to be used in data selection as long as each archive refers to the same sequence and do not have overlapping track names.
Note
Segway does not allow mulitple genomedata archives to contain equivalent tracks names. However if your archives have tracks with matching track names, you may explictly specify to Segway the track names that do not overlap in other genomedata archives and Segway will run as normal.
Tracks
You may specify a subset of tracks using the --track option
which may be repeated. For example:
segway --track dnasei --track h3k36me3
will include the two tracks dnasei and h3k36me3 and no others.
You can run a concatenated segmentation by separating tracks with a comma. For example:
segway --track dnasei.liver,dnasei.blood --track h3k36me3.liver,h3k36me3.blood
Positions
By default, Segway runs analyses on the whole genome. This can be
incredibly time-consuming, especially for training. In reality,
training (and even identification) on a smaller proportion of the
genome is often sufficient. There are also regions as the genome such
as those containing many repetitive sequences, which can cause
artifacts in the training process. The --exclude-coords=file and --include-coords=file options specify BED
files with regions that should be excluded or included respectively.
If both are specified, then inclusions are processed first and the
exclusions are then taken out of the included regions.
Note
BED format uses zero-based half-open coordinates, just like Python. This means that the first nucleotide on chromosome 1 is specified as:
chr1 0 1
The UCSC Genome Browser and Ensembl web interfaces, as well as the wiggle formats use the one-based fully-closed convention, where it is called chr1:1-1.
For example, the ENCODE Data Coordination Center at University of California Santa Cruz keeps the coordinates of the ENCODE pilot regions in this format for `(GRCh37/hg19) http://hgdownload.cse.ucsc.edu/goldenPath/hg19/encodeDCC/referenceSequences/encodePilotRegions.hg19.bed`_
and
`(NCBI36/hg18) http://hgdownload.cse.ucsc.edu/goldenPath/hg18/database/encodeRegions.txt.gz`_.
For human whole-genome studies, these regions have nice
properties since they mark 1 percent of the genome, and were carefully
picked to include a variety of different gene densities, and a number
of more limited studies provide data just for these regions. All
coordinates are in terms of the GRCh37 assembly of the human reference
genome (also called hg19 by UCSC).
After reading in data from a Genomedata archive, and selecting a
smaller subset with --exclude-coords and
--include-coords, the final included regions are referred to
as windows, and are supplied to GMTK for inference. There is no
direction connection between the data in different windows during any
inference process–the windows are treated independently.
An alternative way to speed up training is to use the
--minibatch-fraction=frac option, which will cause Segway to use
a fraction frac or more of genomic positions, chosen randomly at each
training iteration. The --exclude-coords=file and
--include-coords=file options still apply when using minibatch.
The fraction will only apply to the resulting chosen coordinates. For example,
using --minibatch-fraction=0.01 will use a different random one percent of
the genome for each training round. This will allow training to have access to
the whole genome for training while maintaining fast iterations. Using this
option will select on the basis of windows, so the fraction of the genome
chosen will be closer to the specified fraction if the windows are small (but
the chosen fraction will always be at least as large as specified). Therefore,
it is best to combine –minibatch-fraction with –split-sequences. The
likelihood-based training stopping criterion is no longer valid with minibatch
training, so training will always run to –max-train-rounds (100, by default)
if –minibatch-fraction is set.
An alternative way to choose the winning set of parameters
is available through the --validation-fraction=frac or
--validation-coords options. Specifying a fraction frac to
--validation-fraction will cause Segway to choose a fraction frac
or more of genomic positions as a held-out validation set.
--validation-coords=file allows one to explicitly specify genomic
coordinates in a BED-format file, to be used as a validation set. When
using either of these options, Segway will evaluate the model after each
training iteration on the validation set and will choose the winning set
of parameters based on whichever set gives the best validation set likelihood
across all instances.
Note
--exclude-coords is applied to --validation-coords but
--include-coords is not. This allows the user to easily specify
regions of the genome that should not be considered by Segway overall, while
also allowing them to specify a set of validation coordinates in a
straightforward manner.
Resolution
In order to speed up the inference process, you may downsample the
data to a different resolution using the --resolution=res option. This means that Segway will partition the input
observations into fixed windows of size res and perform inference on
the mean of the observation averaged along the fixed window. This can
result in a large speedup at the cost of losing the highest possible
precision. However, if you are considering data that is only generated
at a low resolution to start with, this can be an appealing option.
Warning
You must use the same resolution for both training and identification.
In semi-supervised mode, with the resolution option enabled, the supervision labels are also downsampled to a lower resolution, but by a different method. In particular, segway will partition the input supervision labels into fixed windows of size res and use a modified ‘mode’ to choose which label will represent that window during training. This modified ‘mode’ works according to the following rules:
In general, segway takes the highest-count nonzero label in a given resolution-sized window to be the mode for that window.
In the case of ties, segway takes the lowest nonzero label.
Segway takes the mode to be 0 (no label) if and only if all elements of the window are 0.
Model generation
Segway generates a model (segway.str) and initial parameters
(input.master) appropriate to a dataset using the GMTKL
specification language and the GMTK master parameter file format. Both
of these are described more fully in the GMTK documentation (cite),
and the default structure and starting parameters are described more
fully in the Segway article.
The starting parameters are generated using data from the whole genome, which can be quickly found in the Genomedata archive. Even if you are training on a subset of the genome, this information is not used.
You can tell Segway just to generate these files and not to perform
any inference using the --dry-run option.
Using --num-instances=starts will generate multiple copies of the
input.master file, named input.0.master, input.1.master,
and so on, with different randomly picked initial parameters. Segway
training results can be quite dependent on the initial parameters
selected, so it is a good idea to try more than one. I usually use
–num-instances=10`.
Using --mixture-components will set the number of Gaussian
mixture components per label to use in the model (default 1).
Using --var-floor will set the variance floor for the model,
meaning that if any of the variances of a track falls below this value,
then the variance will be floored (prohibited from falling below
the floor value). This is by default turned off if not using a mixture of
Gaussians; if using a mixture of Gaussians, then it has a default value of
1e-5.
Warning
You should know the scale of your data and set an appropriate variance floor if the scale is very small.
You may substitute your own input.master files but I recommend
starting with a Segway-generated template. This will help avoid some
common pitfalls. In particular, if you are going to perform training
on your model, you must ensure that the input.master file retains
the same #ifdef structure for parameters you wish to train.
Otherwise, the values discovered after one round of training will not
be used in subsequent rounds, or in the annotate or posterior stages.
You can use the --num-labels=labels option to specify the
number of segment labels to use in the model (default 2). You can set
this to a single number or a range with Python slice notation. For
example, --num-labels=5:20:5 will result in 5, 10, and 15 labels
being tried. If you specify --num-instances=starts, then
there will be starts different instances for each of the labels
labels tried.
The question of finding the right number of labels is a difficult one. Mathematical criteria, such as the Bayesian information criterion, would usually suggest using higher numbers of labels. However, the results are difficult for a human to interpret in this case. This is why we usually use ~25 labels for a segmentation of dozens of input tracks. If you use a small number of input tracks you can probably use a smaller number of labels.
There is an experimental --num-sublabels=sublabels
option that enables hierarchical segmentation, where each segment
label is divided into a number of segment sublabels, each one with its
own Gaussian emission parameters. The
output segmentation will be defined according to the
--output-label=output_label option, by default seg,
which will output by (super) segment label as normal. subseg
will output in terms of individual sublabels, only printing out the
sublabel part, and full will print out both the superlabel and
the sublabel, separated by a period. For example, a coordinate
assigned superlabel 1 and sublabel 0 would display as “1.0”.
Using this feature effectively may require manipulation of model
parameters.
Segway allows multiple models of the values of a continuous
observation tracks using three different probability distributions: a
normal distribution (--distribution=norm), a normal distribution
on asinh-transformed data (--distribution=asinh_norm, the
default), or a gamma distribution (--distribution=gamma). For
gamma distributions, Segway generates initial parameters by converting
mean μ and variance σ2 to shape k and scale theta
using the equations μ = kθ and σ2 = kθ2. The ideal methodology for setting gamma parameter
values is less well-understood. I recommend the use of asinh_norm
in most cases.
Virtual Evidence
To use virtual evidence [pearl1988] in Segway, the user specifies a prior probability that a genomic region has a particularly labelled state. In this sense, virtual evidence is a form of semi-supervised learning.
Users supplying virtual evidence while running an entire task together (for
example, train or annotate) will supply the virtual evidence file with
--virtual-evidence. Users supplying virtual evidence while running
steps for a task one at a time will need to specify
--virtual-evidence during the init step in order to properly generate
the input files such as input.master and the triangulation file. During
annotate and posterior a file can be supplied with the
--virtual-evidence option to the init step however it may not be
changed during run.
The virtual evidence file supplied to --virtual-evidence should be of
BED3+2, tab-delimited, format where the 4th column is the label index and the
fifth column is the prior. For example,
chr1 0 1000 0 0.9
will specify a prior probabilty of 0.9 on label 0 for the region of chr1 from 0 to 1000.
If running on multiple concatenated segmentations (worlds), the VE file is in BED3+3 format instead, and the world number must be specified for each row in the last column. If this is omitted and a BED3+2 file is submitted instead, the virtual evidence will be applied to all worlds instead. For example, with two worlds,
chr1 0 1000 0 0.9 0
chr1 0 1000 1 0.05 1
These examples specify prior probabilities over the region of chr1 from 0 to 1000 for both worlds. A prior probability of 0.9 on label 0 for the first world and a prior probability of 0.05 on label 1 for the second world.
At positions for which some labels are given a prior by the user but other labels not, the remaining probability is uniformly distributed amongst the leftover labels. For example, with 4 labels:
chr1 0 1000 0 0.4
all labels but label 0 would be given a prior probability of (1-0.4)/3=0.2.
Pearl, Judea. “Probabilistic reasoning in intelligent systems. 1988.” San Mateo, CA: Kaufmann 23: 33-34.
Model Customization
You can supply your own custom or modified models to Segway by using the
--structure option. The model is defined by the syntax that GMTK
uses. To learn more about using GMTK to create your own models there
the GMTK documentation.
The train and annotate modes of Segway accepts models where states
are given labeled names, rather than using the default GMTK numeric labels.
To assign label names to states of a random variable, the .str structure
file desciption of that variable should contain the line
symboltable: collection("labels_name") where the desired list of label
names is in a name collection in the input.master file named
labels_name. Consult the GMTK documentation for more detailed
information.
These names must contain only alphanumeric characters and will be written to the output annotation file. This feature is not supported for the posterior mode.
Segment duration model
Hard length constraints
The --seg-table=file option allows specification of a
segment table that specifies minimum and maximum segment lengths for
various labels. By default, the minimum segment length is the ruler
length and it is set for all labels. Here is an example of a segment table:
label len
1:4 200:2200:50
0 200::50
4: 200::50
The file is tab-delimited, and the header line with label in one
column and len in another is mandatory. The first column specifies
a label or range of labels to which the constraints apply. In this
column, a range of label values may be specified using Python slice
syntax, so label 1:4 specifies labels 1, 2, and 3. Using 4:
for a label, as in the last row above, means all labels 4 or higher.
The second column specifies three colon-separate values: the minimum
segment length, maximum segment length, and the ruler. In the example
above, for labels 1, 2 and 3, segment lengths between 200 and 2200 are
allowed, with a 50 bp ruler. If either the minimum or maximum lengths
are left unspecified, then no corresponding constraint is applied. If
the ruler is left unspecified the default or set value from the
--ruler-scale option is used.
The ruler is an efficient heuristic that decreases the memory used
during inference at the cost of also decreasing the precision with
which the segment duration model acts. Essentially, it allows the
duration model to switch the behavior of the rest of the model only
after a multiple of scale bp has passed. Note that the ruler must
match all other ruler entries in this file, as well as the option set
with --ruler-scale=scale. (This may become more free
in the future.)
Due to the lack of an epilogue in the model, it is possible to get one segment per sequence that actually does not meet the minimum segment length. This is expected and will be fixed in a future release.
Note that increasing hard minimum or maximum length constraints will greatly increase memory use and run time. You can decrease this performance penalty by increasing ruler size (which makes the precision of the duration model a little coarser), or by using the soft length priors below.
Use these segment lengths along with the supervised learning feature with caution. If you try to create something impossible with your supervision labels, such as defining a 2300-bp region to have label 1, which you have already constrained to have a maximum segment length of 2200, GMTK will produce the dreaded zero clique error and your training run will fail. Don’t do this. In practice, due to the imprecision introduced by the 200-bp ruler, a region labeled in the supervision process with label 1 that is only 2000 bp long may also cause the training process to fail with a zero clique error. If this happens either decrease the size of the ruler, increase the size of the maximum segment length, or decrease the size of the supervision region.
Soft length prior
There is also a way to add a soft prior on the length distribution, which will tend to make the expected segment length 100000, but will still allow data that strongly suggests another length control. The default expected segment length of 100000 can’t be changed at the moment but will in a future version.
You can control the strength of the prior relative to observed
transitions with the --prior-strength=strength option.
Setting --prior-strength=1 means there are as many pseudocounts
due to the prior as the number of nucleotides in the training regions.
The --segtransition-weight-scale=scale option controls
the strength of the prior in another way. It controls the strength of
the length prior relative to the data from the observed tracks. The
default scale of 1 gives the soft transition model equal strength to
a single data track. Using higher or lower values gives comparatively
greater or lesser weight to the probability from the soft length
prior, essentially allowing the prior to have more votes in
determining where a segment boundary is. The impact of the prior will
be a function of both --segtransition-weight-scale and
--prior-strength.
One may effectively use the hard length constraints and soft length priors simultaneously.
Task selection
Segway will perform either (a) model generation and training or (b) identification separately, so it is possible to train on a subset of the genome and annotate on the whole thing. To train, use:
segway train GENOMEDATA TRAINDIR
To annotate, run Segway from the same directory you ran the train task above and specify the TRAINDIR from the results of your training:
segway annotate GENOMEDATA TRAINDIR IDENTIFYDIR
In both cases, replace GENOMEDATA with the Genomedata archive you’re
using. The use of --dry-run will cause Segway to generate
appropriate model and observation files but not to actually perform
any inference or queue any jobs. This can be useful when
troubleshooting a model or task.
Train task
Most users will generate the model at training time, but to specify
your own model there are the --structure=filename and
--input-master=filename options. You can simultaneously
run multiple instances of EM training in parallel, specified with the
--instances=instances option. Each instance consists of
a number of rounds, which are broken down into individual tasks for
each training region. The results from each region for a particular
instance and round are combined in a quick bundle task. It results in
the generation of a parameter file like params.3.params.18 where
3 is the instance index and 18 is the round index. Training for
a particular instance continues until at least one of these criteria is
met:
the likelihood from one round is only a small improvement from the previous round; or
100 rounds have completed.
Specifically, the “small improvement” is defined in terms of the
current likelihood 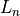 and the log likelihood from the
previous round
and the log likelihood from the
previous round 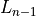 , such that training continues while
, such that training continues while
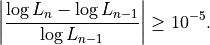
This constant will likely become an option in a future version of Segway.
As EM training produces diminishing returns over time, it is likely that one can obtain acceptably trained parameters well before these criteria are met. Training can be a time-consuming process. You may wish to train only on a subset of your data, as described in Positions.
When all instances are complete, Segway picks the parameter set with the
best likelihood and copies it to params.params.
There are two different modes of training available, unsupervised and semisupervised.
Unsupervised training
By default, Segway trains in unsupervised mode, which is a form of clustering. In this mode, it tries to find recurring patterns suggested by the data without any additional preconceptions of which regions should be tied together.
Semisupervised training
Using the --semisupervised=file option, one can specify
a BED file as a list of regions used as supervision labels. The name
field of the BED File specifies a label to be enforced during
training. For example, with the line:
chr3 400 800 2
one can enforce that these positions will have label 2. You might do this if you had specific reason to believe that these regions were enhancers and wanted to find similar patterns in your data tracks. Using smaller labels first (such as 0) is probably better. Supervision labels are not enforced during the annotate task, and therefore cannot be specified during annotate.
You can also choose to specify a soft assignment for the supervision label. For example, with the line:
chr3 400 800 0:5
one can enforce that these positions will have a label in the range of [0,5). In other words, the label will be restricted to one of {0, 1, 2, 3, 4}. You may want to do this if you know the apparence of the patterns in the regions but you believe they might belong to more than one label. For soft assignment currently we only support a fixed size of the range of labels. For example, you may specify 0:5 and 3:8 in a single supervision label BED file, but you can’t specify 0:5 (range size 5) and 6:8 (range size 2).
To simulate fully supervised training, simply supply supervision labels for the entire training region.
None of the supervision labels can overlap with each other. You should combine any overlapping labels before specifying them to Segway.
It is also possible for nonoverlapping labels to violate the ruler constraints set by Segway for GMTK. This happens when your supervision labels specify a transition that doesn’t fall on a ruler boundary. For example, if your supervision labels are directly adjacent, such as:
chr1 10 20 0:2
chr1 20 30 2:4
and your ruler is set so it won’t allow a transition to occur on 20 then your jobs will terminate with a ‘zero clique’ error. To resolve this, either avoid having directly adjacent superivison labels or, if possible, set –ruler-scale=1 and run Segway again.
General options
The --dont-train=file option specifies a file with a
newline-delimited list of parameters not to train. By default, this
includes the dpmf_always, start_seg, and all GMTK
DeterministicCPT parameters. You are unlikely to use this unless you
are generating your own models manually.
Seeding
Segway can be forced to run with a specified random number generator seed by
setting the SEGWAY_RAND_SEED environment variable. This is an
optional for Segway and is primarily useful for reproducing results future
results or debugging . For example, if you are using bash as your shell you can
run:
SEGWAY_RAND_SEED=1498730685
To set the random number generator seed to the number 1498730685. If you decide to seed the random number generator, it is recommended to pick a number unique for your own usage.
Recovery
Since training can take a long time, this increases the probability
that external factors such as a system failure will cause a training
run to fail before completion. You can use the --recover=dirname option to specify a previous work directory you’re
recovering from.
Annotate task
The annotate mode of Segway uses the Viterbi algorithm to decode the most likely path of segments, given data and a set of parameters, which can come from the train task. Annotate runs considerably more quickly than training. While the underlying inference task is very similar, it must be completed on each region of interest only once rather than hundreds of times as in training.
You must run annotate from the same directory from where the train task was
run. You can either manually set individual input master, parameter, and
structure files, or implicitly use the files generated by the train task
completed in traindir, and referenced in traindir/train.tab. If you
are using training data from an old version of Segway, you must either create a
train.tab file or specify the parameters manually, using
--structure, --input-master, and
--trainable-params.
The --bed=bedfile option specifies where the
segmentation should go. If bedfile ends in .gz, then Segway uses
gzip compression. The default is segway.bed.gz in the working
directory.
You can load the generated BED files into a genome browser. Because the files can be large, I recommend placing them on your web site and supplying the URL to the genome browser rather than uploading the file directly. When using the UCSC genome browser, the bigBed utility may be helpful in speeding access to parts of a segmentation.
The output is in BED format (http://genome.ucsc.edu/FAQ/FAQformat.html#format1), and includes columns for chromosome, start, and end (in zero-based, half-open format), and the label. Other columns to the right are used for display purposes, such as coloring the browser display, and can be safely ignored for further processing. We use colors from ColorBrewer (http://colorbrewer2.org/).
Recovery
The --recover=dirname allows recovery from an
interrupted annotate task. Segway will requeue jobs that never
completed before, skipping any windows that have already completed.
Creating layered output
Segway produces BED files as output with the segment label in the name
field. While this is the most sensible way of interchanging the
segmentation with other programs, it can be difficult to visualize. To
solve this problem, Segway will also produce a layered BED file with
rows for each possible Segment label and thick boxes at the location
of each label. This is what we show in the screenshot figure of the
Segway article. This is much easier to see at low levels of
magnification. The layers are also labeled, removing the need to
distinguish them exclusively by color. While Segway automatically
creates these files at the end of an annotate task, you can also use
segway-layer wit ha standard BED segmentation to repeat the
layering process, which you may want to do if you want to add mnemonic
labels instead of the initial integers used as labels.
segway-layer supports the use of standard input and output
by using - as a filename, following a common Unix convention.
The mnemonic files used by Segway and Segtools have a simple format.
They are tab-delimited files with a header that has the following
columns: old, new, and description. The old column
specifies the original label in the BED file, which is always produced
as an integer by Segway. The new column allows the specification
of a short alphanumeric mnemonic for the label. The description
column is unused by segway-layer, but you can use it to add
helpful annotations for humans examining the list of labels, or to
save label mnemonics you used previously. The row order of the
mnemonic file matters, as the layers will be laid down in a similar
order. Mnemonics sharing the same alphabetical prefix (for example,
A0 and A1) or characters before a period (for example, 0.0
and 0.1) will be rendered with the same color.
segtools-gmtk-parameters in the Segtools package can automatically annotate an initial hierarchical labeling of segmentation parameters. This can be very useful as a first approximation of assigning meaning to segment labels.
A simple mnemonic file appears below:
old new description
0 TSS transcription start site
2 GE gene end
1 D dead zone
Posterior task
The posterior inference task of Segway estimates for each position of
interest the probability that the model has a particular resolution-sized
section given the data. The posterior output does not fall on segment
boundaries since segment duration is itself probabilistically modeled.
This information is delivered in a series of numbered BED files, one for each
segment label. In hierarchical segmentation mode, setting the
–output-label option to full or subseg will cause segway to produce a
wiggle file for each sublabel instead, identified using the label and the
sublabel in the file name before the file extension. For example, the
bedGraph file for label 0, and sublabel 1 would be called
posterior0.1.bedGraph. The individual values will vary from 0 to 100,
showing the percentage probability at each position for the label in that
file. In most positions, the value will be 0 or 100, and substantially
reproduce the Viterbi path determined from the annotate task. The
posterior task uses the same options for specifying a model and
parameters as annotate.
Posterior results can be useful in determining regions of ambiguous labeling or in diagnosing new models. The mostly binary nature of the posterior assignments is a consequence of the design of the default Segway model, and it is possible to design a model that does not have this feature. Doing so is left as an exercise to the reader.
You may find you need to convert the bedGraph files to bigWig format first to allow small portions to be uploaded to a genome browser piecewise.
Recovery
Recovery is not yet supported for the posterior task.
SegRNA
To use SegRNA [segrna2020] with stranded transcriptome datasets, the user provides one track per strand.
Then for each dataset, the user species the tracks for each strand with the --track option separated
by a comma. For example:
segway --track long.polyA.forward,long.polyA.reverse
This command runs Segway in concatenated mode with long.polyA.forward in world 0 and long.polyA.reverse in world 1.
To mimic the direction in which we usually read reverse strand data, from right to left, the user need to set the
--reverse-world option to the world representing reverse strand data. For example:
segway --track long.polyA.forward,long.polyA.reverse --reverse-world=1
Mendez, FANTOM6 consortium, Scott, Hoffman. “Unsupervised analysis of multi-experiment transcriptomic patterns with SegRNA identifies unannotated transcripts” bioRxiv, https://doi.org/10.1101/2020.07.28.225193
Modular interface
Segway additionally supports running the tasks in a more modular manner. Each task is subdivided into 3 common steps:
init: Generates all input files, but does not submit any jobs to GMTK (besides triangulate). At the end of this step the input.master, segway.str, auxillary files, tri files, and the window.bed will all be generated.
run: Submits jobs to GMTK. Produces params and viterbi files.
finish: Selects the best training and identify instances and uses these to generate the output files.
run-round: This step is specific to training. It will run a single round of training and then stop, allowing a user to view results and modify files.
The desired step may be selected by adding it to the task with a hyphen seperating the two, in the form <task>-<step>. For example, to run init for the train step, a user would call segway train-init GENOMEDATA TRAINDIR
Python interface
I have designed Segway such that eventually one may call different components directly from within Python.
You can then call the appropriate module through its main()
function with the same arguments you would use at the command line.
For example:
from segway import run
GENOMEDATA_DIRNAME = "genomedata"
run.main(["train", "--random-starts=3", GENOMEDATA_DIRNAME])
All other interfaces (the ones that do not use a main() function)
to Segway code are undocumented and should not be used. If you do use
them, know that the API may change at any time without notice.
Command-line usage summary
All programs in the Segway distribution will report a brief synopsis
of expected arguments and options when the --help option is
specified and version information when --version is specified.
usage: segway [global_args] COMMAND [args]…
Segmentation and automated genome annotation.
- optional arguments:
- -h, --help
show this help message and exit
- --version
show program’s version number and exit
list of commands:
train create a model with learned parameters - train-init prepare initial models for parallel training - train-run train initial models to completion criterion, in parallel – train-run-round train models for one round, in parallel - train-finish select best model and prepare for annotate
annotate label a genome using a model - annotate-init prepare for parallel annotation - annotate-run annotate the genome, in parallel - annotate-finish concatenate parallel annotation results
posterior infer posterior probabilities of each label across a genome - posterior-init prepare for parallel posterior inference - posterior-run infer posterior probabilities, in parallel - posterior-finish concatenate parallel posterior inference results
Use segway COMMAND –help for help specific to command COMMAND.
- Technical variables:
- -m PROGRESSION, --mem-usage PROGRESSION
try each float in PROGRESSION as the number of gibibytes of memory to allocate in turn (default 2,3,4,6,8,10,12,14,15)
- -S SIZE, --split-sequences SIZE
split up sequences that are larger than SIZE bp (default 2000000)
- -v NUM, --verbosity NUM
show messages with verbosity NUM (default 0)
- --cluster-opt OPT
specify an option to be passed to the cluster manager
- Flags:
- -n, --dry-run
write all files, but do not run any executables
Citation: Hoffman MM, Buske OJ, Wang J, Weng Z, Bilmes J, Noble WS. 2012. Unsupervised pattern discovery in human chromatin structure through genomic segmentation. Nat Methods 9:473-476. https://doi.org/10.1038/nmeth.1937
Utilities
Usage: segway-layer [OPTION]… [INFILE] [OUTFILE]
- Options:
- --version
show program’s version number and exit
- -h, --help
show this help message and exit
- -b FILE, --bigBed=FILE
specify bigBed output file
- -m FILE, --mnemonic-file=FILE
specify tab-delimited input file with mnemonic replacement identifiers for segment labels
- --no-recolor
don’t recolor labels
- -s <ATTR VALUE>, --track-line-set=<ATTR VALUE>
set ATTR to VALUE in track line
Usage: segway-winner [OPTION]… DIR
- Options:
- --version
show program’s version number and exit
- -h, --help
show this help message and exit
- -i, --input-master
print input master file name
- -p, --params
print parameters file name
- -c, --copy
copy files to final winning file locations
- --clobber
overwrite existing files
Environment Variables
- SEGWAY_CLUSTER
Forces segway to use a specific cluster environment. Setting this to ‘local’ forces segway to use run locally and use no cluster environment.
- SEGWAY_NUM_LOCAL_JOBS
Sets the maximum number of jobs when running locally.
- SEGWAY_RAND_SEED
Sets the seed for the random number generator. This is useful for reproducing results.
Running Segway for large jobs
It is highly recommended that a terminal mutliplexer, such as tmux or GNU screen, is used to manage your terminal sessions running Segway. Using either of these programs allows you to create a session to run longer segway jobs that you can safely detatch from without losing your work. For more information, see their respective documentation.
Helpful commands
Here are some short bash scripts or one-liners that are useful:
There used to be a recipe here to continue Segway from an interrupted training run, but this has been replaced by the –old-directory option.
Make a tarball of parameters and models from various directories:
(for DIR in traindir1 traindir2; do
echo $DIR/{auxiliary,params/input.master,params/params.params,segway.str,triangulation}
done) | xargs tar zcvf training.params.tar.gz
Rsync parameters from $REMOTEDIR on $REMOTEHOST to $LOCALDIR:
rsync -rtvz --exclude output --exclude posterior --exclude viterbi \
--exclude observations --exclude "*.observations" --exclude accumulators \
$REMOTEHOST:$REMOTEDIR $LOCALDIR
Print all last likelihoods:
for X in likelihood.*.tab; \
do dc -e "8 k $(tail -n 2 $X | cut -f 1 | xargs echo | sed -e 's/-//g') \
sc sl ll lc - ll / p"; \
done
Recover as much as possible from an incomplete identification run without completing it (which can be done with –old-directory. Note that this does not combine adjacent lines of same segment. BEDTools might be able to do this for you. You will have to create your own header.txt with appropriate track lines.